Asc-Seurat documentation¶
Asc-Seurat (Analytical single-cell Seurat-based web application) is a web application based on Shiny 1. Pronounced as “ask Seurat”, it provides a click-based, easy-to-install, and easy-to-use interface that allows the execution of all steps necessary for scRNA-seq analysis (See Asc-Seurat workflow). It integrates many of the capabilities of the Seurat 2 and Dynverse 3 and also allows an instantaneous functional annotation of genes of interest using BioMart 4.
Asc_seurat relies on multiple R packages. Please, visit the references and check the complete list of packages and their references.
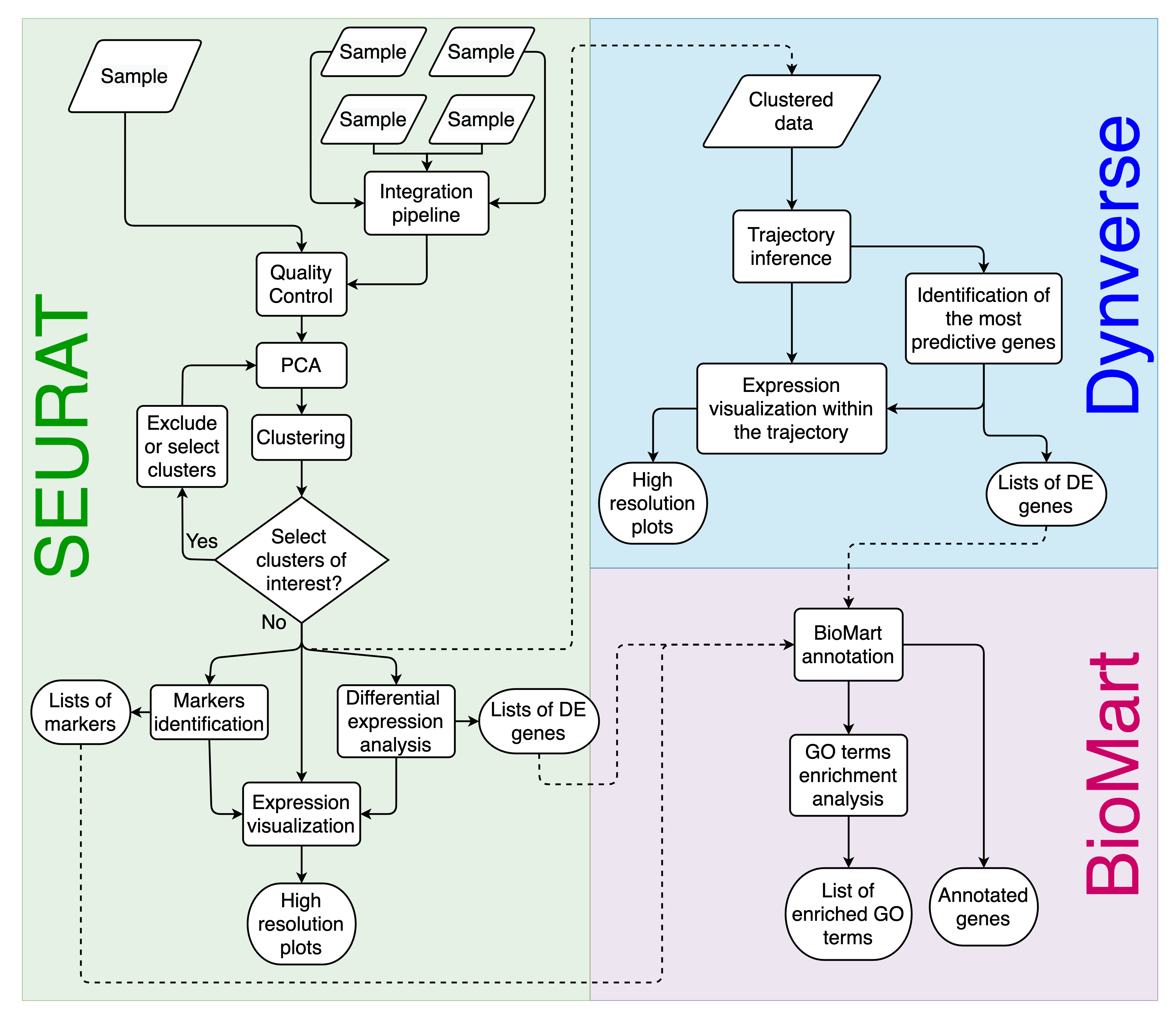
Asc-Seurat workflow overview. Asc-Seurat is built on three analytical cores. Using Seurat, it is possible to explore scRNA-seq data of a population of cells to identify patterns that reflect the cell types of a sample(s) and identify markers and DEGs for each cell type/cluster. By incorporating Dynverse, Asc-Seurat allows the utilization of dozens of models to infer and visualize developmental trajectories (V and VI) and to identify genes differentially expressed on those trajectories (VII). Finally, using BioMart, Asc-Seurat allows immediate functional annotation and GO terms enrichment analysis for many species.¶
Release notes¶
v2.1 - Released on May 26th, 2021.
Changes the assay used for differential expression analysis and visualization to “RNA” when using SCTransform normalization. Therefore, “SCT” assay is used for the steps until clustering the data.
Changes the output of the differential expression analysis to the format required for the visualization tools.
v2.0 - Released on May 19th, 2021.
Inclusion of SCTransform normalization
Addition of stacked violin plots
Addition of multiple-genes dot plot
Improvements on the user interface
Improvements in the app stability
Fix of minor bugs.
v1.0 - Released on March 19th, 2021.
Release of Asc-Seurat.
Reference¶
[1] Pereira WJ, Almeida FM, Balmant KM, Rodriguez DC, Triozzi PM, Schmidt HW, Dervinis C, Pappas Jr. GJ, Kirst M. Asc-Seurat – Analytical single-cell Seurat-based web application. BioRxiv, 2021.
Support Contact¶
Have any questions or suggestions? Please contact us at GitHub.
Footnotes: